OPV1-RPP30DNA
Orthopox virus with human RPP30 DNA control
(RUO). Research Use Only. Not for use in Diagnostic Procedures.
Introduction
The assay OPV1-RPP30DNA is a quantitative polymerase chain reaction (qPCR) test intended for the detection of nucleic acid from several orthopox viruses and detects the conserved protein coding family called “B4R Schlafen-like (Cop-B2R)” (MPXVgp165 Protein ID: gb|URK20603.1). The assay has been experimentally verified to detect a synthetic construct of the Monkeypox virus. Based upon a bioinformatic analysis of sequences, the OPV1 assay should detect most strains of several orthopox viruses including: akhmeta virus, buffalopox, cowpox, ectromelia virus, horsepox, rabbitpox, and vaccinia pox. However, the OPV1 assay will NOT detect variola virus (i.e. causative agent of smallpox) and also likely will not detect racoonpox. This kit is for research use only and should not be used for diagnostic procedures.
The performance of this kit was tested using a standard synthetic DNA construct (orthopox_construct1_DNA) from Twist Biosciences. The limit of detection (LOD) is below 10 copies/reaction.

Tube 1: 20x Primers/Probe specific for Monkeypox and 20x Primers/Probe specific for human RPP30 gene Intron I.
Contents
A mix of primers/probe targeting the orthopox virus is provided in a tube as a 20x concentrated working solution. The fluorophore of the probe is FAM™ (Carboxyfluorescein, a trademark of Life Technologies, Inc) and the quencher is BHQ-1™ (Black Hole Quencher, a trademark of Biosearch Technologies, Inc.) A mix of primers/probe targeting human RPP30 DNA Intron I is also provided in the tube (20x concentrated) as a PCR positive control for human samples. The fluorophore of the probe is HEX™ (Hexachloro-fluorescein, a trademark of Life Technologies, Inc), and the quencher is BHQ-1™.
Kit Handling and Contamination
The kit OPV1-RPP30DNA is shipped at ambient temperature, and should be stored at -30 to -15°C. The kit should be kept on ice once thawed.
Any contamination should be avoided by using appropriate personal protective equipment (PPE), powder free gloves, aerosol barrier pipette tips, and a clean hood.
Experimental
Set up your reaction (20 µL) as follows on ice:
| Component | Volume (µL) |
|---|---|
| InhibiTaq mastermix (2x) | 10 |
| OPV1-RPP30DNA mix (20x) | 1 |
| Sample | 2 |
| Water | 7 |
Note: The composition of this reaction is calculated based on the user manual of InhibiTaq PLUS qPCR Master Mix, from Empirical Biosciences.
A PCR protocol was used in-house for pre-validation on a Bio- Rad CFX96™ Real-Time System, with the following program:
| Step | Thermocycling Protocol: |
|---|---|
| 1 | Incubate @ 95 °C for 2 minutes |
| 2 | Incubate @ 95 °C for 3 seconds |
| 3 | Incubate @ 60 °C for 30 seconds |
| 4 | Plate Read |
| 5 | Go to Step 2, repeat 44x more |
Result Interpretation
After running the qPCR reaction, perform a regression analysis on the data to determine the quantification cycle, Cq. (Cq is preferred over Ct). Each fluorescence channel with a Cq < 38 cycles and final RFU >200 is considered “positive” or “+” in the Table below.
| MPX1 (FAM™) |
RPP30 (HEX™) |
Interpretation and recommendation |
|---|---|---|
| – | – | The PCR reaction failed. Please repeat the experiment |
| – | + | The sample doesn’t contain orthopox DNA |
| + | – | The sample contains orthopox DNA. The sample may not contain human DNA (hRPP30 gene Intron I) |
| + | + | The sample contains orthopox DNA and human DNA (hRPP30 gene Intron I) |
Pre-Validation Experiments
Experiments were performed in triplicate using the experimental procedure given above, but with different samples added to each reaction. The samples used for the validation experiments include water NTC, a 10x diluted human genomic DNA, a water sample spiked with 10000 copies/uL of orthopox_construct1_DNA, and a 10x diluted human genomic DNA spiked with 10000 copies/uL of orthopox_construct1_DNA. The results of these experiments are shown in Figure 1 below:
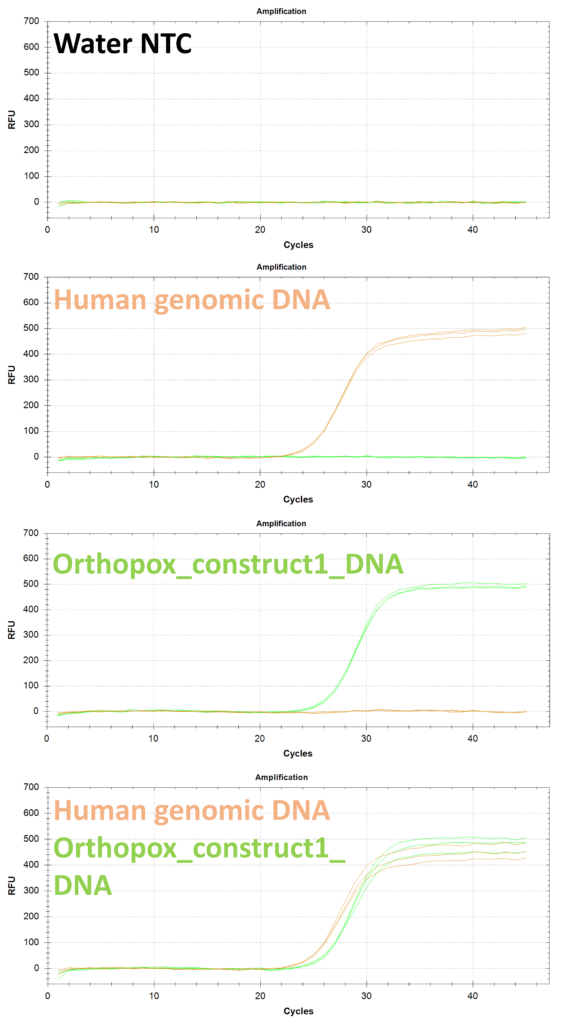
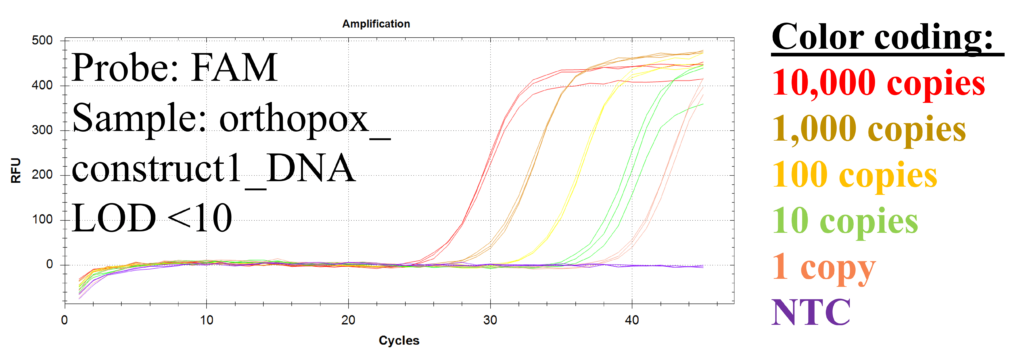
Figure 2: Serial dilution experiments show LOD <10 molecules for orthopox_construct1_DNA.
Conclusion: The data in Figure 1 indicate that the OPV1 primers and probe are compatible with DNAS RPP30DNA positive control primers and probe in a 2-plexing application to detect orthopox in the matrix of human sample extract.
Limit of detection (LOD) was estimated by performing serial dilution experiments in triplicate (Figure 2). For dilution series only the orthopox_construct1_DNA was added. The results show a limit of detection (LOD) <10 copies/reaction.
Contact Us
For assistance, please contact DNA Software using the link: https://www.dnasoftware.com/contact/
Address:
Michigan Life Science and Innovation Center
46701 Commerce Center Dr
Plymouth, MI 48170
Phone: (734) 222-9080