DNAS Completes TMAC Study to Improve PCR Design
DNA Software Completes TMAC Study to Improve PCR Design
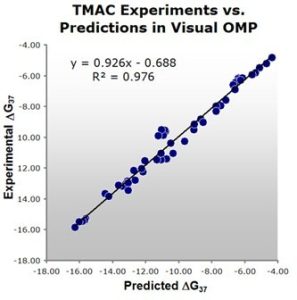
Figure 1: Comparison of experimental versus Visual OMP™ predicted free energies. Note the high correlation of 0.98 between actual experimental results and Visual OMP™ predictions. Fifteen percent of the experimental data in this plot contained the buffer used in bead hybridization technologies marketed by Luminex, Inc.
TMAC Study Summary:
DNA Software is pleased to announce the addition of a free energy correction term for the oligonucleotide duplex stabilizing agent Tetramethylammonium Chloride (TMAC) to its Oligonucleotide Modeling Platform. The equation for this added functionality was derived from experimental UV melting curves of 52 duplexes, having a range of lengths and G+C compositions, in buffers containing TMAC concentrations from 15 mM to 3M TMAC, sodium concentrations ranging from 0.066 to 1.03 M, and magnesium concentrations up to 2 mM.
A marked improvement over the predictive equations from the literature, Visual OMP™ can now predict DGº37 and Tm to within 0.42 kcal/mol and 2.5º C, on average, upon addition of TMAC to the buffers of various PCR, Bead, and Microarray technologies, including those employed by Luminex, Inc.1-4 This added functionality to Visual OMP™ is representative of DNA Software’s commitment to provide our customers with the most current scientific advancements in oligonucleotide modeling and design for the genomic-based assays of the present and the future. This work was supported by NIH SBIR grant # R44 GM076745.
References:
- Diaz, M.R. and Fell, J.W. (2004) “High-Throughput Detection of Pathogenic Yeasts of the Genus Trichosporon”,Journal of Clinical Microbiology, 42, 3696-3706.
- Chevet, E., Lamaitre, G. and Katinka, M.D. (1995) “Low concentrations of tetramethylammonium chloride increase yields and specificity of PCR”, Nucleic Acids Research, 23, 3343-3344.
- Kovarova, M. and Draber, P. (2000) “New specificity and yield enhancer of polymerase chain reactions”, Nucleic Acids Research, 28, e70.
- Wong, C.W., et al. (2007) “Optimization and clinical validation of a pathogen microarray”, Genome Biology, 8, R93.

