DNAS Solves Betaine Thermodynamics for Improved Assay Design
 Betaine Thermodynamics for Improved Assay Design
Betaine Thermodynamics for Improved Assay Design
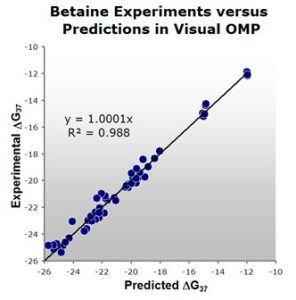
Figure 1: A comparison of experimental versus Visual OMP™ predicted free energies. Note the high correlation of 0.98 between actual experimental results and Visual OMP™ predictions.
Betaine Study Summary:
DNA Software, Inc. is pleased to announce the addition of a free energy correction term for the oligonucleotide duplex hybridization buffer additive betaine monohydrate to its Oligonucleotide Modeling Platform™, including its Visual OMP™ software. The equation for this added functionality was derived from experimental UV melting curves of 63 duplexes, having a range of lengths from 13-24 base pairs with G+C compositions ranging from 23 % to 65 %. The buffers used in this determination contained betaine concentrations from 100 mM to 2.7 M and a variety of sodium and magnesium concentrations.
A marked improvement over the predictive equations from the literature, Visual OMP™ can now predict DGº37 and Tm to within 0.34 kcal/mol and 0.8 ºC, on average, upon addition of betaine to the buffers of various PCR and Microarray technologies.1-6 This added functionality to Visual OMP, in conjunction with that of Tetramethyl-ammonium Chloride (TMAC) released last year, is representative of DNA Software’s commitment to provide its customers with the most current scientific advancements in oligonucleotide modeling and design of genome-based assays. This work was supported by DNA Software’s NIH SBIR grant titled “Database for Modified Nucleotides, Fluorophors and Additives”.
References:
- Ralser, M, Ouerfurth, R., Warnatz, H., Lehrach, H., Yasupo, M. and Krobitsch, S. (2006) “An efficient and economic enhancer mix for PCR”, Biochemical and Biophysical Research Communications, 347, 747-751.
- Wrobel, G, Schlingemann, J., Hummerich, L., Kramer, H., Lichter, P. and Hahn, M. (2003) “Optimization of high-density cDNA-microarray protocols by ‘design of experiments'”, Nuc Acids Res., 31, e67.
- Rees, W.A., Yager, T.D., Korte, J., von Hippel, P. H. (1993) “Betaine Can Eliminate the Base Pair Composition Dependence of DNA Melting”, Biochemistry, 32, 137-144.
- Spink, C. H., Garbett, N. and Chaires, J.B. (2007) “Enthalpies of DNA melting in the presence of osmolytes”, Biophysical Chemistry, 126, 176-185.
- Hong, J., Capp, M.W., Anderson, C.F., Saecker, R. M., Felitsky, D.L., Anderson, M.W. and Record, M.T. (2004) “Preferential Interactions of Glycine Betaine and of Urea with DNA: Implications for DNA Hydration and for Effects of These Solutes on DNA Stability”, Biochemistry, 43, 14744-14758.
- Vasiliskov, V.A., Prokopenko, D.V. and Mirzabekov, A.D. (2001) “Parallel multiplex thermodynamics analysis of coaxial base stacking in DNA duplexes by Oligodeoxyribonucleotide microchips”, Nuc. Acids Res., 29, 2303-2313.

