Primer Dimer Artifacts Are Due to Dimerization
Myth 4: “Primer Dimer” Artifacts Are Due to Dimerization of Primers
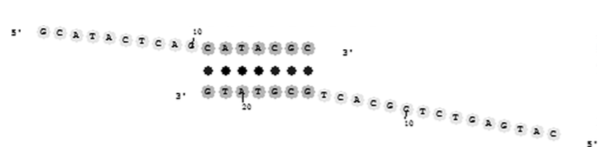
A common artifact in PCR is the amplification of “primer dimers.” The most common conception of the origin of primer dimers is that two primers hybridize at their 3′-ends (see Fig. 8). DNA polymerase can bind to such species and extend the primers in both directions to produce an undesired product with a length that is slightly less than the sum of the lengths of the forward and reverse primers. This mechanism of primer dimerization is certainly feasible and can be experimentally demonstrated by performing thermocycling in the absence of target DNA. This mechanism can also occur when the desired ampli- fication of the target is inefficient (e.g., when one of the primers is designed to bind to a region of the target that is folded into a stable secondary structure). Therefore, most PCR design software packages check candidate primers for 3′-complementarity and redesign one or both of them if the thermodynamic
Fig. 8. Primer dimer hybridized duplex. Note that the 3′-ends of both primers are extensible by DNA polymerase.
stability of the hybrid is above some threshold. Another practical strategy to reduce primer dimer formation is to design the primer to have the last two nucleotides as AA or TT, which reduces the likelihood of a primer dimer structure with a stable hybridized 3′-end (29). For single-target PCR, two primers are present (FP and RP), and there are three different combinations of primer dimers that are possible FP–FP, RP–RP, and FP–RP. For multiplex PCR with N primers, there are NC2 pairwise combinations that are possible, and it becomes harder to redesign the primers so that all of them are mutually compatible. This becomes computationally challenging for large-scale multi- plexing. However, such computer optimization is only partially effective at removing the primer dimer artifacts in real PCRs. Why?
An Alternative Mechanism for Primer Dimer Artifacts
There are some additional observations that provide clues for an alternative mechanism for primer dimerization.
- Generally, homodimers (i.e., dimers involving the same strand) are rarely observed.
- Primer dimer artifacts typically occur at a large threshold cycle number (usually > 35 cycles), which is higher than the threshold cycle number for the desired amplicon.
- Primer dimers increase markedly when heterologous genomic DNA is added.
- Primer dimers are most often observed when one or both of the primers bind inefficiently to the target DNA (e.g., due to secondary structure of the target or weak thermodynamics).
- When the primer dimers are sequenced, there are often a few extra nucleotides of mysterious origin in the center of the dimer amplicon.
Observations 1 and 2 suggest that DNA polymerase does not efficiently bind to or extend primer duplexes with complementary 3′-ends. Observation 2 could also be interpreted as meaning that the concentration of the primer duplex is quite low compared with the normal primer-target duplex. In the early stages of PCR, however, observations 3 and 5 suggest that background genomic DNA may play a role in the mechanism of primer dimer formation. Observation 4 suggests that primer dimerization needs to occur in the early rounds of PCR to prevent the desired amplicon from taking over the reactions in the test tube. Figure 9 illustrates a mechanism that involves the genomic DNA in the early cycles of PCR and that provides an explanation for all five observations.
The mechanism presented in Fig. 9 can also be checked for by computer, but searching for such a site in a large genome can be quite computationally demanding. The ThermoBLAST algorithm developed by DNA Software, Inc. can meet this challenge (see myth 5).
Additional Concerns for Primer Dimers
Two primers can sometimes hybridize using the 5′ end or middle of the sequences. Such structures are not efficiently extensible by DNA polymerase. Such 5′-end primer hybrids, however, can in principle affect the overall equilibrium for hybridization, but generally, this is a negligible effect that is easily minimized by primer design software (i.e., if a primer is predicted to form a significant interaction with one of the other primers, then one or both of the primers are redesigned to bind to a shifted location on the target). If a polymerase is used that has exonuclease activity (e.g., Pfu polymerase), then it is possible that hybridized structures that would normally be non-extensible might be chewed back by the exonuclease and create an extensible structure. Indeed, it is observed that PCRs done with enzymes that have exonuclease
Fig. 9. Genomic DNA can participate in the creation of both the desired amplicon and the primer dimerization artifact. Notice that despite the presence of a few mismatches, denoted by “x,” the middle and 5′-ends of the primers are able to bind to the target stronger than they would bind to another primer molecule. Note that this mechanism does not require very strong 3′-end complementarity of the primers P1 and P2. Instead, this mechanism requires that sites for P1 and P2 are close to each other.
activity have a much higher incidence of primer dimer formation and mishybridization artifacts. Thus, for PCR, “proofreading” activity can actually be harmful.