Author: wahost
DNA Software, Inc. Completes Assay Design and Optimization Training
Assay Design and Optimization Training
Ann Arbor, MI – August 20th, 2009 – DNA Software, Inc. just completed a successful, two day training of assay design at its new offices in downtown Ann Arbor, MI.
 The first day of training on assay design focused on the science of nucleic acid thermodynamics, basic and advanced function in Visual OMP™, advanced assay design including multiplex PCR and microarrays, as well as high-throughput and scripting applications for DNA Software’s Developer Edition™ products.
The first day of training on assay design focused on the science of nucleic acid thermodynamics, basic and advanced function in Visual OMP™, advanced assay design including multiplex PCR and microarrays, as well as high-throughput and scripting applications for DNA Software’s Developer Edition™ products.
Each class participants received a free trial license to Visual OMP™ with the ThermoBLAST™ and Modifieds™ modules and performed several hands-on case studies on assay development.
The second day of training included an overview of DNA Software’s newly released Modifieds™ module. This new add-on feature for Visual OMP™ allows users to simulate and develop assays that incorporate modified nucleotides. It contains thermodynamic parameters for LNA, PNA, Morpholino, DeoxyU, Inosine, Iso-C, and Iso-G in the context of DNA and RNA. The ThermoBLAST™ module, algorithm, and science were also reviewed during the second day. ThermoBLAST™ represents a much more accurate method than BLAST to check your assay designs against whole genomes to ensure they do not mishybridize to unintended targets. As with previous training classes, part of the class was dedicated to customers’ specific research projects.
Given the success of this class DNA Software is proposing a class in Amsterdam in mid-October and another one in Ann Arbor in later November. If you are interested in joining us for either of these training sessions, please contact:sales@dnasoftware.com or call +1 734 222 9080.
About DNA Software, Inc.
DNA Software, Inc. was founded in 2000 to commercialize the advances in nucleic acid chemistry discovered by world-renowned expert Dr. John SantaLucia, Jr. (the company’s Chief Scientific Officer). DNA Software’s technologies have become the standard of excellence for nucleic acid research and diagnostics development.
DNA Software is a unique software company considering that it conducts original wet lab research. The company’s software programs are the most accurate and comprehensive tools available because their advanced algorithms and models are driven by a large database of thousands of diverse, wet lab-derived, experimental results. DNA Software’s tools correctly design and simulate complicated experiments on the first attempt. Customers can simulate thousands of experiments with the software before running a single experiment in a wet lab. Thus, DNA Software’s technologies save customers significant time, resources, and money that would have been wasted on trial-and-error experimentation.
DNA Software offers contract research, custom software development, commercial web applications, and scientific consulting for nucleic acid research. The company also licenses packaged software tools that help scientists quickly and accurately develop new assays, diagnostics, and therapeutics. OMP™ (Oligonucleotide Modeling Platform™), the core engine that runs all of DNA Software’s tools, models in silico the folding and hybridization of nucleic acids with exceptional accuracy. Visual OMP™ is a platform program for the design, simulation, and analysis of probes/primers, RT-PCR, qPCR, Multiplex PCR, Taqman, Beacons, Scorpions, Microarrays, SNP detection, FRET assays, RNAi, LATE-PCR, plus new formats. OMP Developer EditionTM (OMP DE™) brings customizable, command line, high-throughput computing power to large assay design projects or enterprise applications. ThermoBLAST™ quickly and accurately scans DNA and RNA against genome databases, identifies all off-target hybridizations, and tells you if these mishybridized probes/primers are extensible by polymerase. Modifieds™ allows you to simulate and develop assays that incorporate modified nucleotides (e.g. LNA, PNA, Morpholino, DeoxyU, Inosine, Iso-C, and Iso-G).
The company has recently expanded its original DNA and RNA-based, molecular biology solutions to include modified nucleotides, oligonucleotide kinetics, and structural biology.
We Awarded NIH Funding to Develop Nucleic Acid Technologies
NIH Funding to Develop Nucleic Acid Technologies
ANN ARBOR, MI – July 22, 2009 – DNA Software, Inc. has been awarded three Fast Track SBIR grants from the National Institutes of Health (NIH Funding) to develop original in silico technologies to predict 3D structures of RNA-based molecules, improve diagnostics via modified nucleotides, and model the reaction rates of DNA and RNA experiments. The company successfully completed its milestones for Phase I of each project and recently began work on Phase II.
DNA Software, Inc. is a unique life sciences technology company that conducts original wet lab research, develops advanced bioinformatics tools, sells molecular biology software, offers structural biology services, and provides advanced scientific consulting. DNA Software’s products and services improve the work of scientists who conduct nucleic acid-based research and latest NIH funding will give a big boost to that type of research.
Nucleic acids, commonly known as DNA (Deoxyribonucleic acid) and RNA (Ribonucleic acid), are the molecular building blocks of life, form the basis of genetic blueprints, and direct activities in living organisms. DNA Software’s tools help scientists to quickly and accurately develop new medical solutions from this nucleic acid information.
In Silico 3D Structure Prediction to Improve and Accelerate Drug Discovery
The genomics era has produced a flood of interesting macromolecular sequences, but there is a shortage of corresponding 3-dimensional (3D) structural information. As a result, there is a lack of structural understanding of the mechanism of the vast majority of known nucleic acid sequences. Half of all clinically used antibiotics target bacterial ribosomes, which are RNA-based molecules. Yet, the 3D structures for the complete ribosomes of many pathogenic bacteria are still unknown. Thus, the ribosome represents an important target for structural prediction and future drug development, including antibiotics for pathogenic bacteria that have developed drug resistance.
DNA Software has a breakthrough technology, called Nucleic Acid CAD™ (NA-CAD™), which expands nucleic acid sequence information into all-atom 3D structure predictions of diverse RNA-based structures with near-crystal structure accuracy. With this NIH project, DNA Software is modeling ribosome structure and function and aims to predict the 3D structures of complete ribosomes of clinically relevant pathogens.
Novel Software for Modeling PCR Reaction Rates and Diagnostics with Modified Nucleotides
While current software programs can model the thermodynamics (i.e. the results at equilibrium after a long time) of DNA and RNA reactions, none can model the kinetics (i.e. how fast the reaction takes place). In addition, most software can only model DNA or RNA. None can accurately model the large variety of modified nucleotides that are added to or replace DNA and RNA sequences to increase the sensitivity and specificity of nucleic acid-based diagnostic tests and therapeutics. DNA Software is combining the unique features of kinetics and modified nucleotides with its existing OMP™ (Oligonucleotide Modeling Platform™) software to create a first-rate tool to develop ultra-fast, sensitive, and selective diagnostics. OMP™ has become the gold standard for assay development. It is utilized by prominent research organizations from around the world and across many industries (e.g. biotech, pharmaceutical, diagnostics, biodefense, reagents, government, academic, and institutes).
DNA Software recently released its first software product for modified nucleotides. The Modifieds™ software module allows users to simulate and design assays that incorporate modified nucleotides within the OMP™ program. Modifieds™ currently includes LNA, PNA, DeoxyU, Morpholino, Inosine, Iso-C, and Iso-G. DNA Software is currently deriving additional thermodynamic parameters for many more modified nucleotides in its wet lab and this NIH funding will help us develop more tools for research. These parameters will be released to each customer as they become available and as part of the Modifieds™ software license. Modifieds™ is a unique product on the market. Leading research organizations, including the Centers for Disease Control and Prevention, are utilizing Modifieds™ to improve their assay development.
About DNA Software, Inc.
DNA Software, Inc. was founded in 2000 to commercialize the advances in nucleic acid chemistry discovered by world-renowned expert Dr. John SantaLucia, Jr. (the company’s Chief Scientific Officer). DNA Software’s technologies have become the standard of excellence for nucleic acid research and diagnostics development.
DNA Software is a unique software company considering that it conducts original wet lab research. The company’s software programs are the most accurate and comprehensive tools available because their advanced algorithms and models are driven by a large database of thousands of diverse, wet lab-derived, experimental results. DNA Software’s tools correctly design and simulate complicated experiments on the first attempt. Customers can simulate thousands of experiments with the software before running a single experiment in a wet lab. Thus, DNA Software’s technologies save customers significant time, resources, and money that would have been wasted on trial-and-error experimentation.
DNA Software offers contract research, custom software development, commercial web applications, and scientific consulting for nucleic acid research. The company also licenses packaged software tools that help scientists quickly and accurately develop new assays, diagnostics, and therapeutics. OMP™ (Oligonucleotide Modeling Platform™), the core engine that runs all of DNA Software’s tools, models in silico the folding and hybridization of nucleic acids with exception accuracy. Visual OMP™ is a platform program for the design, simulation, and analysis of probes/primers, RT-PCR, qPCR, Multiplex PCR, Taqman, Beacons, Scorpions, Microarrays, SNP detection, FRET assays, RNAi, LATE-PCR, plus new formats. OMP Developer EditionTM (OMP DE™) brings customizable, command line, high-throughput computing power to large assay design projects or enterprise applications. ThermoBLAST™ quickly and accurately scans DNA and RNA against genome databases, identifies all off-target hybridizations, and tells you if these mishybridized probes/primers are extensible by polymerase. Modifieds™ allows you to simulate and develop assays that incorporate modified nucleotides (e.g. LNA, PNA, Morpholino, DeoxyU, Inosine, Iso-C, and Iso-G).
The company has recently expanded its original DNA and RNA-based, molecular biology solutions to include modified nucleotides, oligonucleotide kinetics, and structural biology.
DNA Software, Inc. Honored as 2009 “Michigan 50 Companies to Watch”
Michigan 50 Companies to Watch
ANN ARBOR, MI – April 29, 2009 – DNA Software, Inc. has been recognized as one of the 2009 “Michigan 50 Companies to Watch,” an awards program sponsored by the Edward Lowe Foundation and presented by Michigan Celebrates Small Business.
DNA Software, Inc. is a unique life sciences technology company that conducts original wet lab research, develops advanced bioinformatics tools, sells molecular biology software, and provides advanced scientific consulting. DNA Software’s products and services improve the work of scientists who conduct nucleic acid-based research.
Nucleic acids, commonly known as DNA (Deoxyribonucleic acid) and RNA (Ribonucleic acid), are the molecular building blocks of life, form the basis of genetic blueprints, and direct activities in living organisms. DNA Software’s tools help scientists to quickly and accurately develop new medical solutions from this nucleic acid information.
Award Underscores DNA Software’s Innovation, Impact, and Growth
“The Michigan 50 Companies to Watch award is an appreciated honor that recognizes DNA Software’s ongoing efforts to conduct original science, develop and commercialize innovative technologies, help the scientific community solve major challenges, and expand our business in new areas”, states Donald Hicks, President and CEO of DNA Software, Inc.
Mr. Hicks continues “DNA Software has a successful record of turning NIH SBIR funded R&D into commercial products. Many research organizations around the world and a variety of industries have benefited from DNA Software’s technologies. DNA Software is helping the US government to address important health challenges. For example, the Center for Disease Control and Prevention uses DNA Software’s Visual OMP™ assay development software to design life-saving diagnostic tests. The U.S. Army Medical Research Institute of Infectious Diseases is utilizing DNA Software’s OMP Developer Edition™ high-throughput, computer cluster software to anticipate and mitigate biological threats. In addition, DNA Software has partnered with the Department of Homeland Security to expand the use of DNA Software’s ThermoBLAST ™ genome scanning software for biodefense applications.”
Jeff Machak, Vice President of Business Development, adds “Since its founding in 2000, DNA Software has steadily grown its commercial business and built an extensive customer list. Over the past few years, we have refocused our business model and experienced growth in revenue through increased sales, SBIR funding, and government contract work. Moreover, we have expanded our strategic alliances with government agencies, academic institutions, Fortune 500 companies, and international sales partners. DNA Software has doubled its staff within the past year and hired top talent from Pfizer, Johnson & Johnson, and Siemens. Most recently, we have expanded our R&D pipeline to create exciting new technologies for improved diagnostics development and drug discovery.”
About the “50 Companies to Watch” Award
“In today’s economy, Companies to Watch has become a real game changer,” says Penny Lewandowski, director of entrepreneurship development at the Edward Lowe Foundation, a nonprofit operating foundation based in Michigan. “It changes the conversation from slow-growth, layoffs and closings, to companies that are expanding, hiring, and moving beyond traditional markets.”
Companies nominated for the Michigan 50 Companies to Watch list must be second-stage companies and meet criteria for number of employees, sales revenue, and growth. In addition, the companies must be privately-held and headquartered in Michigan.
Winners were selected by judges from the banking, economic development, entrepreneurship development, and venture capital communities. Judges evaluated the nominees’ intent and capacity to grow based on factors which included exceptional entrepreneurial leadership, competitive advantage, and demonstrable growth in sales and employee base.
Information about the 2009 Michigan 50 Companies to Watch program can be found athttp://Michigan.CompaniesToWatch.org
For information about Michigan Celebrates Small Business, visit http://MichiganCelebrates.biz
About DNA Software, Inc.
DNA Software, Inc. was founded in 2000 to commercialize the advances in nucleic acid chemistry discovered by world-renowned expert Dr. John SantaLucia, Jr. (the company’s Chief Scientist). DNA Software’s technologies have become the standard of excellence for nucleic acid research and diagnostics development.
DNA Software is a unique software company considering that it conducts original wet-lab research. The company’s software programs are the most accurate and comprehensive tools available because their advanced algorithms and models are driven by a large database of thousands of diverse, wet lab-derived, experimental results. DNA Software’s tools correctly design and simulate complicated experiments on the first attempt. Customers can simulate thousands of experiments with the software before running a single experiment in a wet lab. Thus, DNA Software’s technologies save customers significant time, resources, and money that would have been wasted on trial-and-error experimentation.
DNA Software offers contract research, custom software development, commercial web applications, and scientific consulting for nucleic acid research. The company also licenses packaged software tools that help scientists quickly and accurately develop new assays, diagnostics, and therapeutics. OMP™ (Oligonucleotide Modeling Platform™), the core engine that runs all of DNA Software’s tools, models in silico the folding and hybridization of nucleic acids with exception accuracy. Visual OMP™ is a platform program for the design, simulation, and analysis of probes/primers, RT-PCR, qPCR, Multiplex PCR, Taqman, Beacons, Scorpions, Microarrays, SNP detection, FRET assays, RNAi, LATE-PCR, plus new formats. OMP Developer EditionTM (OMP DE™) brings customizable, command line, high-throughput computing power to large assay design projects or enterprise applications. ThermoBLAST™ quickly and accurately scans DNA, RNA, or hybrid sequences against genome databases and eliminates all assay false positives or probe/primer mishybridizations.
The company has recently expanded its original DNA and RNA-based, molecular biology solutions to include modified nucleotides, oligonucleotide kinetics, and structural biology.
DNAS Solves Betaine Thermodynamics for Improved Assay Design
 Betaine Thermodynamics for Improved Assay Design
Betaine Thermodynamics for Improved Assay Design
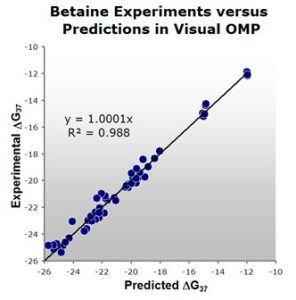
Figure 1: A comparison of experimental versus Visual OMP™ predicted free energies. Note the high correlation of 0.98 between actual experimental results and Visual OMP™ predictions.
Betaine Study Summary:
DNA Software, Inc. is pleased to announce the addition of a free energy correction term for the oligonucleotide duplex hybridization buffer additive betaine monohydrate to its Oligonucleotide Modeling Platform™, including its Visual OMP™ software. The equation for this added functionality was derived from experimental UV melting curves of 63 duplexes, having a range of lengths from 13-24 base pairs with G+C compositions ranging from 23 % to 65 %. The buffers used in this determination contained betaine concentrations from 100 mM to 2.7 M and a variety of sodium and magnesium concentrations.
A marked improvement over the predictive equations from the literature, Visual OMP™ can now predict DGº37 and Tm to within 0.34 kcal/mol and 0.8 ºC, on average, upon addition of betaine to the buffers of various PCR and Microarray technologies.1-6 This added functionality to Visual OMP, in conjunction with that of Tetramethyl-ammonium Chloride (TMAC) released last year, is representative of DNA Software’s commitment to provide its customers with the most current scientific advancements in oligonucleotide modeling and design of genome-based assays. This work was supported by DNA Software’s NIH SBIR grant titled “Database for Modified Nucleotides, Fluorophors and Additives”.
References:
- Ralser, M, Ouerfurth, R., Warnatz, H., Lehrach, H., Yasupo, M. and Krobitsch, S. (2006) “An efficient and economic enhancer mix for PCR”, Biochemical and Biophysical Research Communications, 347, 747-751.
- Wrobel, G, Schlingemann, J., Hummerich, L., Kramer, H., Lichter, P. and Hahn, M. (2003) “Optimization of high-density cDNA-microarray protocols by ‘design of experiments'”, Nuc Acids Res., 31, e67.
- Rees, W.A., Yager, T.D., Korte, J., von Hippel, P. H. (1993) “Betaine Can Eliminate the Base Pair Composition Dependence of DNA Melting”, Biochemistry, 32, 137-144.
- Spink, C. H., Garbett, N. and Chaires, J.B. (2007) “Enthalpies of DNA melting in the presence of osmolytes”, Biophysical Chemistry, 126, 176-185.
- Hong, J., Capp, M.W., Anderson, C.F., Saecker, R. M., Felitsky, D.L., Anderson, M.W. and Record, M.T. (2004) “Preferential Interactions of Glycine Betaine and of Urea with DNA: Implications for DNA Hydration and for Effects of These Solutes on DNA Stability”, Biochemistry, 43, 14744-14758.
- Vasiliskov, V.A., Prokopenko, D.V. and Mirzabekov, A.D. (2001) “Parallel multiplex thermodynamics analysis of coaxial base stacking in DNA duplexes by Oligodeoxyribonucleotide microchips”, Nuc. Acids Res., 29, 2303-2313.
DNAS Exhibits Its Expanding Technology Portfolio
Ann Arbor, Mich. – October 17, 2007 – DNA Software, Inc. exhibited its science and software developments and growing products and services portfolio at the annual MichBio Expo in Lansing, MI.
DNA Software received The Michigan Investment and Commercialization Success Award in 2003. DNA Software is among the select-few, award-winning, start-up companies that successfully commercialized its technology, continues to expand its R&D and commercial offerings, and is working toward diversifying and growing Michigan’s struggling economy.
DNA Software continues to explore opportunities to develop the biotech industry within Michigan. The company networks with local business-accelerator organizations such as the Michigan Economic Development Corporation (MEDC), MichBio, SPARK, and BioArbor. DNA Software has collaborated with, licensed technology from, licensed software to, and provided consulting services to Michigan universities. When Pfizer Inc decided to close its Ann Arbor R&D facility in early 2007, DNA Software attracted, hired, and retained local talent. Moreover, DNA Software has consulting relationships with former Pfizer scientists and continues to seek expertise in the areas of computational chemistry, structural biology, and mathematical modeling. Interested scientists are encouraged to contact DNA Software.
About DNA Software, Inc.
DNA Software is the leading provider of innovative software and consulting solutions for nucleic acid research. DNA Software combines award-winning science with cutting-edge software technology and offers the industry’s most comprehensive, automated, and predictive software for designing accurate DNA and RNA hybridization assays and reducing costly false positives and negatives.
DNAS Completes TMAC Study to Improve PCR Design
DNA Software Completes TMAC Study to Improve PCR Design
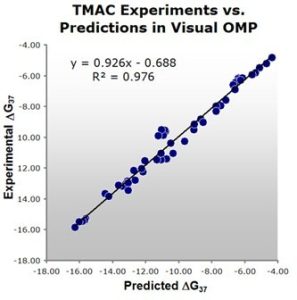
Figure 1: Comparison of experimental versus Visual OMP™ predicted free energies. Note the high correlation of 0.98 between actual experimental results and Visual OMP™ predictions. Fifteen percent of the experimental data in this plot contained the buffer used in bead hybridization technologies marketed by Luminex, Inc.
TMAC Study Summary:
DNA Software is pleased to announce the addition of a free energy correction term for the oligonucleotide duplex stabilizing agent Tetramethylammonium Chloride (TMAC) to its Oligonucleotide Modeling Platform. The equation for this added functionality was derived from experimental UV melting curves of 52 duplexes, having a range of lengths and G+C compositions, in buffers containing TMAC concentrations from 15 mM to 3M TMAC, sodium concentrations ranging from 0.066 to 1.03 M, and magnesium concentrations up to 2 mM.
A marked improvement over the predictive equations from the literature, Visual OMP™ can now predict DGº37 and Tm to within 0.42 kcal/mol and 2.5º C, on average, upon addition of TMAC to the buffers of various PCR, Bead, and Microarray technologies, including those employed by Luminex, Inc.1-4 This added functionality to Visual OMP™ is representative of DNA Software’s commitment to provide our customers with the most current scientific advancements in oligonucleotide modeling and design for the genomic-based assays of the present and the future. This work was supported by NIH SBIR grant # R44 GM076745.
References:
- Diaz, M.R. and Fell, J.W. (2004) “High-Throughput Detection of Pathogenic Yeasts of the Genus Trichosporon”,Journal of Clinical Microbiology, 42, 3696-3706.
- Chevet, E., Lamaitre, G. and Katinka, M.D. (1995) “Low concentrations of tetramethylammonium chloride increase yields and specificity of PCR”, Nucleic Acids Research, 23, 3343-3344.
- Kovarova, M. and Draber, P. (2000) “New specificity and yield enhancer of polymerase chain reactions”, Nucleic Acids Research, 28, e70.
- Wong, C.W., et al. (2007) “Optimization and clinical validation of a pathogen microarray”, Genome Biology, 8, R93.
DNA Software Awarded NIH Grant to Improve Probe, Primer Specificity
DNA Software Awarded NIH Grant to Improve Probe and Primer Specificity
Ann Arbor, Michigan – September 29, 2006 – DNA Software has been awarded a fast track grant by the National Institute of Health (NIH) to develop a cutting-edge software prototype, called ThermoBLAST™.
ThermoBLAST™ is a revolutionary new tool that combines the accuracy of DNA Software’s proprietary OMP Nearest Neighbor model with the speed and breadth of the BLAST algorithm to produce a fast yet exhaustive method for genome scale searches. ThermoBLAST™ will be integrated into DNA Software’s OMP™ (Oligonucleotide Modeling Platform™) software resulting in automated design of more specific probes and primers that can be designed more quickly and more economically. ThermoBLAST™ will allow scientists to develop better assays, while saving time and money.
ThermoBLAST™ will make a major positive contribution to the work of public and private researchers worldwide. Scientists who are designing DNA tests to identify or discover the function of important, disease related genes will be able to “test their tests” in a computer, without having to go through the lengthy lab testing process for each assay. The most difficult DNA-based tests are also the most important. There are very small differences between the genetic code of deadly pathogens and harmless microorganisms. ThermoBLAST™ will greatly improve the ability of scientists to design DNA probes that yield consistent conclusive results, and avoid costly false negatives and positives.
This fast track grant, entitled “Development of ThermoBLAST: Improving the Specificity of Probes and Primers,” is scheduled from September 29, 2006 to April 30, 2008.
About DNA Software, Inc.
DNA Software, Inc. combines science and software to enable industrial genomics through advances in technologies based on nucleic acids. The company’s first software platform, OMP™ (Oligonucleotide Modeling Platform™), models in silico the folding and hybridization of single-stranded nucleic acids with great accuracy. The company combines OMP™ with scientific consulting, custom software development, and custom laboratory research to deliver state-of-the-art support for designing and developing of nucleic acid based technologies.
DNAS Awarded NIH Grant to Study Modified Nucleic Acids
Ann Arbor, Michigan – September 7, 2006 – DNA Software has been selected for a Grant issued by the National Institute of Health (NIH). The fast track grant, entitled “Database for Modified Nucleotides, Fluorophores and Additives” has been approved for Phase I and Phase II funding. Phase I, scheduled from November 1, 2005 to May 1, 2006, has recently been completed. Phase II is scheduled to start October 1, 2006.
Phase I
In Phase I of this project survey measurements on morpholino modified oligonucleotides and on 5-methylisocytosine and isoguanine match and mismatch duplexes have been performed. These measurements form the basis for the design of some of the experiments for phase II.
Previously determined effects of glycerol on nucleic acid denaturation have been incorporated into Visual OMP™. The last phase I accomplishment was to add previously determined parameters for deoxyinosine substitutions in DNA duplexes to Visual OMP™; these will be available to Visual OMP™ users in the next update.
Phase II
Besides completing the morpholino:RNA match, Morpholino:RNA dangling end, and iC and iG thermodynamic parameter databases in phase II, systematic measurements will be made to also complete the thermodynamic parameter database for PNA. These measurements include PNA:RNA matches, PNA:PNA matches, PNA hairpins, PNA:DNA mismatches and PNA:DNA dangling ends.
The other Phase II goals include completing the thermodynamic parameter database for 18 commonly used labels (fluorophores, quenchers, and biotin), and determining the thermodynamic corrections for duplex stability as a function of concentration for the additives DMSO, Betaine, SYBR-Green, TMAC, and formamide.
Lastly, these newly determined parameters and dependencies will be incorporated into Visual OMP™, and a module to design custom probes with modified nucleotides will be developed.
DNA Software expects Phase II to be completed by October of 2007. Astrid Tuin, DNA Software’s lab director states that “The Phase II experiments, analysis of the data and software development are carefully planned, so we expect phase II of this exciting project to go smoothly. All of the goals should be accomplished on time.”
Background and significance of the project
Well designed morpholino oligonucleotides can be used as an efficient and specific oligonucleotide-based antisense reagent in the field of functional genomics to match biological function(s) with gene sequence.
The pairing capability of iC and iG has been employed for molecular recognition and site-specific incorporation by DNA polymerase. In complex assays where high levels of natural DNA of known or unknown sequences exist, the orthogonal pairing capability of iC-iG allows for specific molecular recognition to take place without interference.
PNA has excellent properties, like chemical and biological stability, enhanced hybridization affinity to DNA and RNA, and incorporation of non-natural nucleobases, that can be used for diagnostics, antigene and antisense therapies, and antiviral and antimicrobial PNA therapies.
By determining the thermodynamic influences of labels (fluorophores, quenchers) and different additives, Visual OMP™ can help improve design of (multiplex) Real Time PCR assays, leading to higher efficacy and specificity.
About DNA Software, Inc.
DNA Software, Inc. combines science and software to enable industrial genomics through advances in technologies based on nucleic acids. The company’s first software platform, OMP™ (Oligonucleotide Modeling Platform™), models in silico the folding and hybridization of single-stranded nucleic acids with great accuracy. The company combines OMP™ with scientific consulting, custom software development, and custom laboratory research to deliver state-of-the-art support for designing and developing of nucleic acid based technologies.
DNAS attends American Society of Human Genetics Annual Meeting.
Ann Arbor, Mich. DNA Software is attending the American Society of Human Genetics Annual Meeting on October 25-29, 2005. This year’s ASHG Annual Meeting is taking place in Salt Lake City, Utah. Several of DNA Software’s senior engineers will be available to answer your questions.
This year’s theme of american society of human genetics meeting is “Realizing the Promise of the Human Genome Project“, which as the ASHG’s president states “..it is relevant to clinicians trying to diagnose and characterize birth defects and developmental disorders; to counselors informing and supporting patients and families about these disorders; to basic scientists trying to decipher the pathogenesis of Mendelian disorders; to geneticists trying to dissect contributions to complex traits; and to all of us trying to translate the wealth of genomic information into function.”
About DNA Software, Inc.
DNA Software, Inc. combines science and software to enable industrial genomics through advances in technologies based on nucleic acids. The company’s first software platform, OMP™ (Oligonucleotide Modeling Platform™), models in silico the folding and hybridization of single-stranded nucleic acids with great accuracy. The company combines OMP™ with scientific consulting, custom software development, and custom laboratory research to deliver state-of-the-art support for designing and developing of nucleic acid based technologies.
DNAS Launches Visual OMP 3 Simulating DNA, RNA based Assays
Genomic researchers can now design more specific and sensitive nucleic acid experiments and reduce trial and error bench work with DNAS Visual OMP 3.
Ann Arbor, Mich. – July 10, 2003 – DNAS Launches Visual OMP™ 3 for Designing and Simulating DNA and RNA based Assays. DNA Software, a leading provider of software, scientific consulting and lab research for genomic researchers, announced today the launch of Visual OMP™ 3. This product is the latest advancement of the company’s proprietary Oligonucleotide Modeling Platform™ (OMP™) which provides researchers with an in silico laboratory environment for designing PCR and microarray based experiments. With Visual OMP™ 3, genomic researchers are able to design more sensitive and specific assays and reduce trial and error bench work.
“DNAS Visual OMP™ 3 provides researchers with the ability to automatically design multiple sets of primers and probes that are optimized for specific experiments yet reduce cross-hybridization and mis-hybridization between oligos and targets,” stated Don Hicks, COO and vice president of DNA Software.
Unlike existing software products, DNA Software’s proprietary ThermoBlast algorithms evaluate the thermodynamic energy associated with every possible competitive reaction and predicts the concentration each will form in a reaction. Probes and primers can be tailored to emphasize specificity, sensitivity, melting temperature or length. DNAS Visual OMP™ 3 is the outcome of nearly ten years research by Dr. John SantaLucia, chief science officer of DNA Software and associate professor at Wayne State University.
“We’ve been using OMP to solve real-world assay design problems on a consulting basis for over two years. Visual OMP™ 3 brings together all that we’ve learned about using in silico technology to design better assays with less effort,” noted Mark Kielb, president of DNA Software.
Visual OMP™ 3’s integrated folding engine allows scientists the ability to visualize target and oligo structure much faster than other available products and includes the first sequence dependent hairpin model. Visual OMP™ 3’s in silico laboratory is further enhanced by integration of the third-party tools BLAST and ClustalW.
DNA Software also offers a OMP Developer Edition™ (OMP DE™), that allows programmers to integrate OMP™ into high-throughput applications. OMP DE™ provides an application programming interface (API) for scripting OMP™ access and linking it with existing databases and supports client/server, distributed (GRID), and network installation.
DNA Software’s Visual OMP™ 3 is available for the Windows operating environment. OMP DE™ is available for Windows, Linux, UNIX, and Alpha operating systems.
Academic pricing is available for all products. For more information or to schedule a demonstration, please contact DNA Software at +1 (734) 222-9092.
About DNA Software, Inc.
DNA Software, Inc. combines science and software to enable industrial genomics through advances in technologies based on nucleic acids. The company’s first software platform, OMP™ (Oligonucleotide Modeling Platform™) models in silico the folding and hybridization of single-stranded nucleic acids with great accuracy. The company combines OMP™ with scientific consulting, custom software development, and custom laboratory research to deliver state-of-the-art support for designing and developing of nucleic acid based technologies.




 Betaine Thermodynamics for Improved Assay Design
Betaine Thermodynamics for Improved Assay Design